Computational biology and bioinformatics
Computational Biology and Bioinformatics is the application and development of computational methods to solve important problems in biology and biotechnology. At USJ, we develop novel machine learning models to learn the hidden patterns from biological data (DNA and proteins) and correlate them with the observed physiochemical properties and biological activities of these molecules. These models help us predict and design molecules with improved functions for drug development. In addition, our research involves modelling and simulating three-dimensional structures of proteins, ligands, membranes, and their assembly to gain insights into their interactions and functions.
Computer-Aided Drug Discovery
Principal Investigator: Shirley W. I. Siu
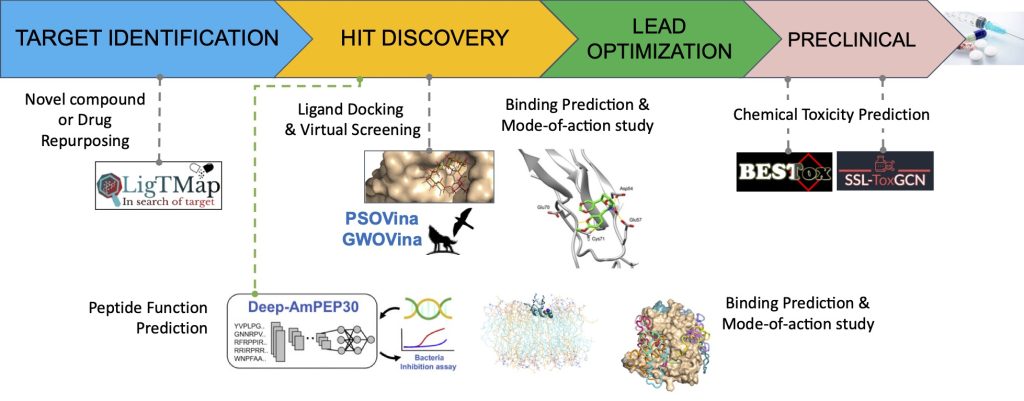
Our research projects cover the early stages of drug discovery, from target identification to hit discovery and drug property evaluation. We have worked on both small molecules and short peptides as drug candidates, and have developed computational tools for novel drug discovery.
Antimicrobial Peptides
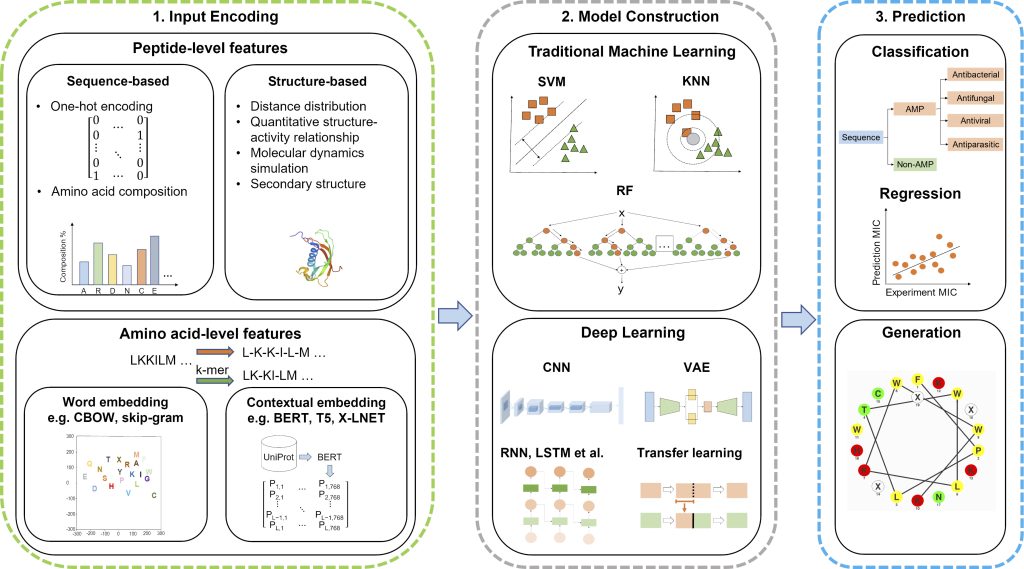
Antimicrobial peptides (AMPs) are a valuable source of antimicrobial agents and a potential solution to the problem of multidrug resistance. In particular, short-length AMPs have been shown to have enhanced antimicrobial activity, higher stability, and lower toxicity to human cells. To support the discovery and development of effective AMPs, we are developing machine learning and deep learning methods to recognize amino acid patterns of AMPs using publicly available data. Although relatively good classification can be achieved, it is still challenging to accurately predict the antibacterial assay value of peptides. The following figure summarizes the machine learning and deep learning approaches to predict AMPs.
For more information, see our review paper Recent Progress in the Discovery and Design of Antimicrobial Peptides using Traditional Machine Learning and Deep Learning, Antibiotics 2022.
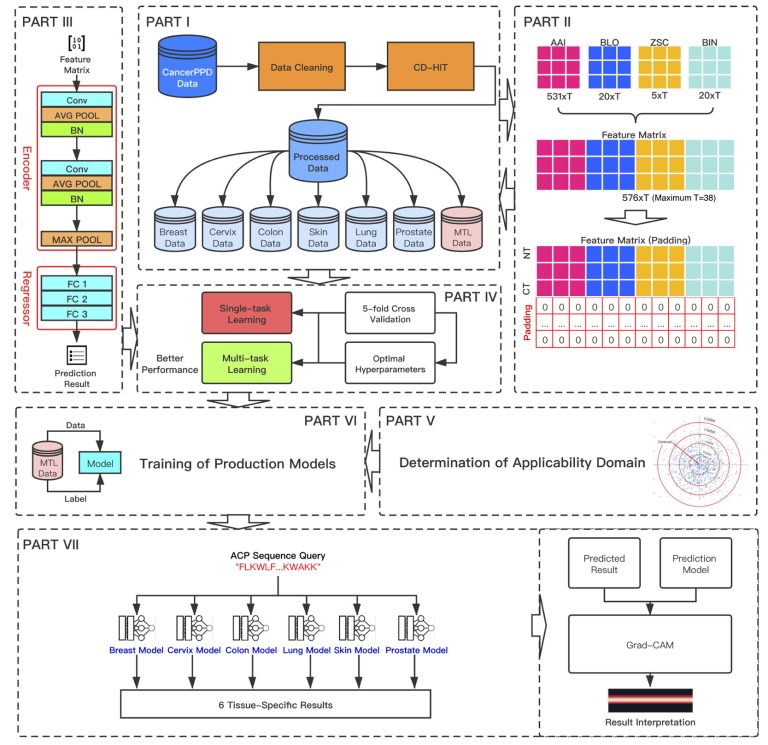
Anticancer Peptides